|
1 |
python pyinstxtractor.py file_to_decompile |
.NET Bio is Significantly Faster on .Net Core 2.0
The .NET Bio library contains libraries for genomic data processing tasks like parsing, alignment, etc. that are too computationally intense to be undertaken with interpreted languages like Python or R, but can be efficiently performed in code that is typed and compiled, like Java and C#, without suffering the productivity burden of coding in C/C++.
Historically, .NET Bio ran on two different frameworks. On windows one could leverage all the substantial engineering Microsoft invested in their CLR to create fast and stable programs. On Mac or Linux, one could use the independently implemented Mono Framework, which was a solid effort by a significantly less resourced team that performed well, but never really came close to the quality of the Microsoft implementation. In practice, as bioinformaticians are weened on the Linux/GCC toolchain and infrastructure, Mono was the frequently used option. This led to performance and reliability issues.
All of this changed when Microsoft decided to open source .NET , releasing a version of their framework for Mac, Linux and Windows called .NET Core. This was a watershed moment, as it allows open source writers to take advantage of all the advanced memory management and compilation techniques Microsoft developed. Better Garbage collection, advanced pre-compilation and vector instructions are all allowing higher level languages like C# to perform nearly as well as aggressively optimized C/C++ for many practical applications.
To test out the new possibilites, I compared the time it takes to parse all the reads in a 1.2 GB BAM file using either the old Mono runtime or the new dotnet core runtime on my MacBook computer. The test code was simple counting of reads after they were deserialized from the compressed format and converted into objects.
|
1 2 3 4 5 |
var p = new BAMParser(); int count = 0; foreach (var read in p.Parse(fname)) { count += 1;} Console.WriteLine(count); |
htslib, a C library for the same task that uses tons of bit operations, and is about as performant as one can get 1)This is for methods that fully parse the BAM read, PySAM, which lazily decodes the data, can finish a simple counting task in about 15 seconds, but this isn’t representative of a workflow which would require full decoding..
|
1 2 3 4 5 6 |
# Run example on .net core 2.0 time dotnet test.dll # On Mono time mono test.dll # Using samtools samtools view test.bam | wc -l |
Profiling Each Platform
Mono: The Mono platform allows us to easily profile the code using instruments on Mac OSX. Examining the profile of the running code, we can see where the bottlenecks are. A portion of time is spent inlibz doing decompression of the underlying gzipped data, but it appears that much is also spent on garbage collection associated events, e.g. preparing for memory allocation with mono_handle_new.

.NET Core 2.0: Being newly available on multiple platforms, there is no good way to profile .NET Core 2.0 on Mac OSX. Although typical profilers cannot understand the C# JITted code, we can look at the compiled C/C++ code and here we see that .NET is spending twice as much time in libz as a percentage than Mono is. Since much of this task is simply decompressing data, more time in the decompression library means the rest of the code is running faster, and is a good sign. 
To further profile the managed code, I wrote a custom MacOS Profiler that changed how each method was compiled to insert a hook that allowed me to track the enter and exit time of each function with a P/Invoke call to C++ code at the entrance and exit of each managed method frame, and so calculated the inclusive time spent in each method. Although the addition of this instrumentation is expected to greatly skew the results, we saw that the majority of the managed code is spent parsing the data, and much of that is spent doing validations!
| Method Name | Inclusive Time (s) | Call Count |
|---|---|---|
| Bio.IO.BAM.BAMParser.GetAlignedSequence | 2.01235 | 65255 |
| Bio.Util.Helper.StringContainsIllegalCharacters | 0.67041 | 1487805 |
| Bio.IO.SAM.SAMOptionalField.set_Tag | 0.551263 | 711275 |
| Bio.IO.SAM.SAMOptionalField.set_VType | 0.549131 | 711275 |
| Bio.IO.SAM.SAMOptionalField.IsValidTag | 0.547594 | 711275 |
| Bio.IO.SAM.SAMOptionalField.IsValidVType | 0.54615 | 711275 |
| Bio.IO.SAM.SAMOptionalField.set_Value | 0.543942 | 711275 |
| Bio.IO.SAM.SAMOptionalField.IsValidValue | 0.543379 | 711275 |
libz, samtools spends half of it’s time there, and the other half basically doing all the decompression operations to convert the binary data into usable data (in sam_format1). SAMTools is optimized C code with minimal error checking, so is a rough upper bound on the speed that could be obtained.

Conclusions
The C# code was written in a manner free of performance concerns and has tons of validation checks and unnecessary copies. Yet .NET core performed only 2X slower than high performance C code. This is due to the fact that thanks to the engineering in .NET core, the code can now run twice as fast on Mac OSX as it used to.Although C# is still slower than C, it is the clear winner here. The SAMTools code bottlenecks at a method that although heavily optimized is horrifically difficult to read. In contrast, the .NET code bottlenecks at a method that is filled with nearly gratuitous memory allocations, and a plethora of checks where useful exceptions could be thrown and meaningful error messages delivered if assumptions aren’t met. Plus, it has all the advantages of memory safety and quality tooling we’ve come to expect in 2017. It’s intriguing to ponder how the benchmark would look if the C# code had been written in a style as performance based as the C code, by avoiding unnecessary allocations, using pointers, or using the new ref returns available in C# 7.0. I’ve shown previously such changes s could significantly increase the parsers performance.
The .NET ecosystem is moving along well for meaningful scientific computing. Importantly, although time-spent C/C++ code will always be more performant, as .NET Core is now open source, we can embed any time-critical functionality in the runtime itself now, allowing us to have things like ludicrously performant suffix-arrays interacting with parsers. .NET Core 2.0 is a big step forward.
References
| 1. | ↑ | This is for methods that fully parse the BAM read, PySAM, which lazily decodes the data, can finish a simple counting task in about 15 seconds, but this isn’t representative of a workflow which would require full decoding. |
Why R Math Functions on Windows are Slow, and How to Fix It
The log Function on MinGW / Windows
Comparing the assembly code of the log function as generated with MinGW on windows with the assembly generated for the log function on Mac OSX shows why the code is slow. Below is the assembly code for log function as decompiled from MinGW (their implementation is basically a cut/paste from GNU libc). The crucial feature here is that most instructions start with an “f” indicating that they are using the floating point registers, and there is one instruction thefyl2xp1 that is one of the most expensive operations out there. This instruction takes a log using hardware, and is known to be slower than most other possible ways to calculate log for modern machines.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 |
0000000000000010 <__logl_internal>: 10: d9 ed fldln2 12: db 2a fldt (%rdx) 14: d9 c0 fld %st(0) 16: dc 25 e4 ff ff ff fsubl -0x1c(%rip) # 0 <one> 1c: d9 c0 fld %st(0) 1e: d9 e1 fabs 20: dc 1d e2 ff ff ff fcompl -0x1e(%rip) # 8 <limit> 26: df e0 fnstsw %ax 28: 80 e4 45 and $0x45,%ah 2b: 74 12 je 3f <__logl_internal+0x2f> 2d: dd d9 fstp %st(1) 2f: d9 f9 fyl2xp1 31: 48 89 c8 mov %rcx,%rax 34: 48 c7 41 08 00 00 00 movq $0x0,0x8(%rcx) 3b: 00 3c: db 39 fstpt (%rcx) 3e: c3 retq 3f: dd d8 fstp %st(0) 41: d9 f1 fyl2x 43: 48 89 c8 mov %rcx,%rax 46: 48 c7 41 08 00 00 00 movq $0x0,0x8(%rcx) 4d: 00 4e: db 39 fstpt (%rcx) |
The log Function on Mac OSX
Demonstrating faster assembly code is the Mac OSX implementation of log shown below. The key feature here is that it uses XMM registers and has a more modern and performant implementation. Note that many other implementations (e.g. the Microsoft compiler) also use faster versions like this. That is, Windows is not slow, R is slow on Windows. It’s all the same Intel chips under the hood.|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 |
+0x00 vmovq %xmm0, %rax +0x05 shrq $32, %rax +0x09 movl %eax, %edx +0x0b subl $1048576, %eax +0x10 cmpl $2145386496, %eax +0x15 jae "0x7fff9793e100+0xe4" +0x1b andl $1048575, %eax +0x20 addl $4096, %eax +0x25 andl $2088960, %eax +0x2a shrl $9, %eax +0x2d addl $3222802432, %edx +0x33 sarl $20, %edx +0x36 vcvtsi2sdl %edx, %xmm1, %xmm1 +0x3a vmovsd 112942(%rip), %xmm6 +0x42 vandpd %xmm6, %xmm0, %xmm0 +0x46 vmovsd 74890(%rip), %xmm7 +0x4e vorpd %xmm7, %xmm0, %xmm0 +0x52 leaq 112999(%rip), %rdx +0x59 vmovss (%rdx,%rax), %xmm3 +0x5e vpsllq $32, %xmm3, %xmm3 +0x63 vfmsub213sd %xmm7, %xmm0, %xmm3 +0x68 vmovddup %xmm3, %xmm3 +0x6c vaddpd -48(%rdx), %xmm3, %xmm4 +0x71 vfmadd213pd -32(%rdx), %xmm3, %xmm4 +0x77 vmovapd -16(%rdx), %xmm5 +0x7c vaddsd %xmm3, %xmm5, %xmm5 +0x80 vmulpd %xmm3, %xmm5, %xmm5 +0x84 vmulpd %xmm5, %xmm4, %xmm4 +0x88 vmovhlps %xmm4, %xmm5, %xmm5 +0x8c vmulsd %xmm5, %xmm4, %xmm4 +0x90 je "0x7fff9793e100+0xc7" +0x92 vmovss 4(%rdx,%rax), %xmm5 +0x98 vpsllq $32, %xmm5, %xmm5 +0x9d vmulsd 112867(%rip), %xmm1, %xmm0 +0xa5 vmulsd 112851(%rip), %xmm1, %xmm1 +0xad vaddsd 8(%rdx,%rax), %xmm0, %xmm0 +0xb3 vaddsd %xmm5, %xmm1, %xmm1 +0xb7 vaddsd %xmm4, %xmm0, %xmm0 +0xbb vaddsd %xmm3, %xmm0, %xmm0 +0xbf vaddsd %xmm1, %xmm0, %xmm0 +0xc3 vzeroupper +0xc6 retq +0xc7 vmovss 4(%rdx,%rax), %xmm5 +0xcd vpsllq $32, %xmm5, %xmm5 +0xd2 vaddsd %xmm5, %xmm3, %xmm3 +0xd6 vaddsd 8(%rdx,%rax), %xmm4, %xmm0 +0xdc vaddsd %xmm3, %xmm0, %xmm0 +0xe0 vzeroupper +0xe3 retq |
Solving the problem
Ideally we could just push a faster log implementation to the R core repo for Windows, but the fact of the matter is the R core team does not accept code changes that only improve performance (and fair play to them, R is the most cross platform software out their, and that wasn’t easy to do). Also, realistically the current implementation is feasible for most peoples work. So the solution is to switch out the log function for a better one only when it matters, and if it matters that means most of the library is already written in compiled code. Where to get a cross-platform compatible log function? It seems the folks over at Julia ran into a similar problem with slow Log on different machines. They looked at two version in libm (e.g. Libm Version #1 and Libm Version #2), and also some really crazy implementations such as this one available from Yepp. However, what they settled on is a very good function, and we can port those over directly into R. An example is shown in the pull request here.Profiling Rcpp package code on Windows
Change compilation settings to add in symbol settings
A default R installation typically has certain compiler settings placed in the equivalent of the C:\Program Files\R\R-3.3.1\etc\x64\Makeconf that strips information needed for profiling during the Rcpp compilation process, in particular a line which reads: DLLFLAGS=-s . To override this and add some additionally needed flags, one should add a folder and file to their home directory which overrides and appends necessesary compilation flags. To a file located at a location equivalent to C:\Users\YOURNAME\.R\Makevars on your machine (note the ‘.’ before R), add the following lines:|
1 2 |
CXXFLAGS+=-gdwarf-2 DLLFLAGS= |
-gdwarf-2 appears in the compilation messages, and that -s is missing in the final linker step.
Run a profiler which understands MinGW compiled code
The next key step is to run a profiler which can understand the Unix like symbols on windows. Two free and good options are Very Sleepy and AMD’s code analyst (which also works on Intel chips). Very Sleepy is very good at basic timings and providing stack traces, while AMD’s profiler is able to drill down to the assembly of a process. Both profilers are good but an example with AMD is shown below.- Open the program and setup a quick session to start and run a sample R script that uses your code, such as in the example shown below.

- Next run the profiler and get ready to look at results. For example, here I can see that half the time was spent in my code, versus half in the R core’s code (generating random numbers)
 And digging further down I can see at the assembly level what the biggest bottlenecks were in my code
And digging further down I can see at the assembly level what the biggest bottlenecks were in my code
 Its often helpful to look at the original source files in addition to the assembly, and this can be enabled by setting directory information by Tools-> CodeAnalyst Options -> Directories.
Its often helpful to look at the original source files in addition to the assembly, and this can be enabled by setting directory information by Tools-> CodeAnalyst Options -> Directories.
C# vs. Java, Xamarin vs. Oracle, Performance Comparison version 2.0
Today I noticed the SIMD implementation of the Mandelbrot set algorithm I blogged about last year was successfully submitted to the language shootout webpage. However, I was a bit disappointed to see the C# version was still slower than the Java version, despite my use of the special SIMD instructions (though I was pleased that a quick port of my code to F# absolutely annihilates the OCAML version of this benchmark).
I had benchmarked my code as faster than the Java code on my Mac, the CentOS cluster at my work and two Ubuntu virtual machines (Mono >3.0, LLVM compiler). What gave?
Undeterred, I thought I would try to use SIMD to improve another C# benchmark, and took a shot at the N-body benchmark test. Once again, the version I wrote, in my hands, was much faster. But when submitted to the shoot-out and accepted, it was either much slower, or didn’t even run. My submission using the SSE instructions not only didn’t beat Java, it was actually slower than the original C# version!
Below are the timings on my top-of-the-line Mac for the two benchmarks, in both cases we see that the C# program runs in 80-90% of the time the Java Program takes. There are several key take aways here.
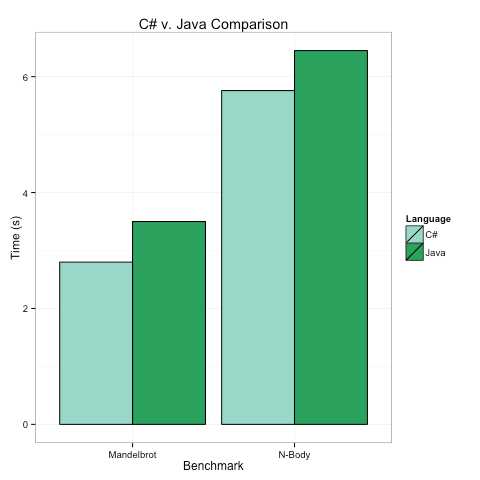
C# and Java have no meaningful performance differences.
C# may use a lot less memory, but this is my optimized C# code and it is only beating the optimized Java by <= 20%. When does that matter?C# and Java’s similar performance has different underpinnings
There are a lot of ways code can be fast, and I was surprised that Java and C# achieve similar speeds in very different ways.
The key advantage of the C# code was the SIMD instructions, which in theory give a ~2X speed up. However, they only win by ~20%. Why?
I think the assembly gives the answer. Just some quick snippets tell the whole story. Here is some C# code for the N-Body problem compiled to assembly:
|
1 2 3 4 5 6 7 8 |
00000034 vsubpd 0x18(%esi), %xmm2, %xmm2 00000039 vmulpd %xmm1, %xmm1, %xmm3 0000003d vmulpd %xmm2, %xmm2, %xmm4 00000041 vhaddpd %xmm3, %xmm3, %xmm3 00000045 vhaddpd %xmm4, %xmm4, %xmm4 00000049 vaddpd %xmm4, %xmm3, %xmm3 0000004d vsqrtpd %xmm3, %xmm4 00000051 vmulpd %xmm4, %xmm3, %xmm3 |
The important insights into performance from the assembly here are:
- Similar instructions are stacked on top of each other, allowing for pipelining (e.g. the same vhaddpd instruction follows the same vhaddpd, and both use different registers, so can execute simultaneously).
- The “p” in the instructions (i.e. vhadd-“p”-d). This stands for “packed” meaning we are packing/doing two operations at once via SIMD.
- Only registers XMM1-XMM4 appear in the instructions. There are more registers available, and more pipelining possible, but the Mono/LLVM compiler appears to only use the low-number/scratch registers. It is easier to write compilers that obey this restriction.
Now let’s compare that to some Java assembly emitted by the Oracle runtime for the same benchmark:
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
0x0000000109ce2315: vmulsd %xmm11,%xmm1,%xmm2 0x0000000109ce231a: vmulsd %xmm11,%xmm9,%xmm3 0x0000000109ce231f: vmulsd %xmm9,%xmm9,%xmm0 0x0000000109ce2324: vmulsd %xmm1,%xmm1,%xmm5 0x0000000109ce2328: vmovsd 0x38(%r12,%rdx,8),%xmm4 0x0000000109ce232f: vaddsd %xmm0,%xmm5,%xmm5 0x0000000109ce2333: vmovsd 0x30(%r12,%rdx,8),%xmm6 0x0000000109ce233a: vmovsd 0x28(%r12,%rdx,8),%xmm7 0x0000000109ce2341: vmovsd 0x20(%r12,%rbp,8),%xmm0 0x0000000109ce2348: vmovsd 0x40(%r12,%rbp,8),%xmm8 0x0000000109ce234f: vsubsd %xmm0,%xmm12,%xmm0 0x0000000109ce2353: vmulsd %xmm8,%xmm9,%xmm9 0x0000000109ce2358: vmulsd %xmm11,%xmm0,%xmm14 0x0000000109ce235d: vmulsd %xmm8,%xmm0,%xmm15 0x0000000109ce2362: vmulsd %xmm0,%xmm0,%xmm0 0x0000000109ce2366: vmulsd %xmm8,%xmm1,%xmm1 |
The important insights here are:
- Java uses the version of the instructions with “s” (i.e. vmul-“s”-d) meaning single, and gets no SSE advantage. No doubt this makes writing compilers easier.
- However, Java is using lots of registers, XMM15 shows up!
- As a result of using all the registers used, the Java code has great pipelining. Note that up to 5 vmulsd instructions show up at once. The JVM is simply brilliant at packing operations together, and this means that even though I, the programmer, was smart and used SSE2, my C# code only won by 20%, and not 200%. It’s hard to beat Java pipelining.
All of which makes me wonder. What if we took the high-level language advantages of C# (low-overhead value types, easy interop via pointers, baked in generics, etc.). And combined them with the low-level advantages of the JVM (better array bounds check elimination, pipelining compiler optimizations, and maybe someday the occasional stack allocation of reference type variables…)
NuMTs, mtDNA sequencing and Aligners
|
1 |
bwa mem hg19.fna f.fq r.fq > simulatedData.sam |
 For reads that did align to the nucleus, the MAPQ was typically 0, but could be as high as 60 and had an unexplained peak at 27.
For reads that did align to the nucleus, the MAPQ was typically 0, but could be as high as 60 and had an unexplained peak at 27.
 And below shows the normalized coverage by positions across the mtDNA, clearly some regions are more affected by NuMTs.
And below shows the normalized coverage by positions across the mtDNA, clearly some regions are more affected by NuMTs.
 Reads from the first and last 500 bp of the mtDNA are poorly aligned by bwa. It appears most go to chromosome 17, but their true location is
belied by their mate pair. In fact only 0.6% of reads in this region that
map to the nuclear DNA do not have their paired read map to the mtDNA.
Reads from the first and last 500 bp of the mtDNA are poorly aligned by bwa. It appears most go to chromosome 17, but their true location is
belied by their mate pair. In fact only 0.6% of reads in this region that
map to the nuclear DNA do not have their paired read map to the mtDNA.
 I also wanted to see how reads that represent a heteroplasmic deletion would be handled. I simulated reads that either spanned or included a deletion randomly chosen to be in the mtDNA, again virtually all mapped to the mitochondria, and the coverage profile looked similar to the simulation with complete reads. Perhaps most reassuringly, almost all reads are mapped. Checking for unmapped reads with the command:
I also wanted to see how reads that represent a heteroplasmic deletion would be handled. I simulated reads that either spanned or included a deletion randomly chosen to be in the mtDNA, again virtually all mapped to the mitochondria, and the coverage profile looked similar to the simulation with complete reads. Perhaps most reassuringly, almost all reads are mapped. Checking for unmapped reads with the command:
|
1 |
samtools view -f 4 Simulated.bam |
The .NET Bio BAM Parser is Smoking Fast
 The comparison to samtools is based on the system time to parse the file using this task on the
same unix server used for the C# tests.
The comparison to samtools is based on the system time to parse the file using this task on the
same unix server used for the C# tests.
samtools view Test.bam | wc -l
And is just meant to give a general idea of the performance comparison, as there are several important differences between the samtools and .NET Bio test. The C# version was not allowed to reuse memory for objects as it was supposed to be working as a data producer, while the Samtools version processes reads one at a time and does reuse memory. C# also made a lot of dictionaries to aid quick access to the read groups, which isn’t done by samtools. However, samtools had to write the files to the output pipe, while the C# version did not, which undoubtably introduces a fair bit of overhead for it. Both tools however, are clearly plenty fast and at this stage further performance improvements would come from lazy evaluation (or not sticking unnecessary information like the original quality scores in the BAM files!), and the language won’t matter much.
Performance Comments
One task when parsing BAMs is unpacking lots of information that is packed together in arrays. In SAMtools and the current .NET Bio parser, this is done with lots of unpacking of bits by integer manipulations. For example this code from SAMTools:
|
1 2 3 4 5 6 7 8 9 10 11 |
uint32_t x[8], block_len = data_len + BAM_CORE_SIZE, y; int i; assert(BAM_CORE_SIZE == 32); x[0] = c->tid; x[1] = c->pos; x[2] = (uint32_t)c->bin<qual<l_qname; x[3] = (uint32_t)c->flag<n_cigar; x[4] = c->l_qseq; x[5] = c->mtid; x[6] = c->mpos; x[7] = c->isize; |
|
1 2 3 |
AlignmentData ad; fixed (byte* alblck = alignmentBlock) { ad = *((AlignmentData*)alblck); } |
Using Selectome with .NET Bio, F# and R
Example Walk Through.
Step 0: Setup F#–If you haven’t used F# before, you can download Tsunami. Open the program and add references to Bio.Selectome using the #r command and open the relevant namespaces. (highlight and hit Alt+Enter to run in the console).
|
1 2 3 4 5 6 7 8 |
#r @"C:\Users\Nigel\Software\Bio.dll" #r @"C:\Users\Nigel\Software\Bio.Pamsam.dll" #r @"C:\Users\Nigel\Software\Bio.Selectome.dll" open System open System.IO; open Bio.Selectome open System.Linq |
Step 1: Make Gene List – Selectome requires ensembl identifiers be used in queries.To create a set of interesting genes, I first downloaded the full set of genes from the MitoCarta website. These genes are identified by Entrez IDs, while selectome uses Ensembl IDs, so to convert between these I used the GeneID converter website to create a flatfile of the new ids. Given this flatfile, we can load it and convert it to typed classes as follows:
|
1 2 3 4 |
let fname= @"C:\SomeFolder\FullListOfGenes.csv" let data=File.ReadAllLines fname |> Array.map (fun x->x.Split(',')) type Gene = {Gene: string; EntrezID:string; EnsemblID:string} let genesForQuery=Array.map (fun (x:string[])-> {Gene=x.[4];EnsemblID=x.[3];EntrezID="Not Set"}) data |
Step 2: Query Selectome – All selectome data is accessed through the
SelectomeDataFetcher class. This class will return a SelelctomeQueryResult that will let you know if the query was successful. Currently, the queries will only be successful for genes that exist in the database and have data available for the full vertebrate tree. If no data is available the Result will be NoResultsFound, if selectome returned data but there was no tree available for all vertebrates(but maybe just primates) the result will be NoVeterbrateTreeDataFound. We want to extract genes from query results that successfully returned data for the vertebrate tree.
|
1 2 3 4 |
let getGene (gene:Gene) : SelectomeQueryResult = SelectomeDataFetcher.FetchGeneByEnsemblID(gene.EnsemblID) let queriedData = genesForQuery |> Array.map getGene |> Array.filter (fun x->x.Result=QUERY_RESULT.Success) |> Array.map (fun x-> x.Gene) |
Step 3: Determine how many genes show positive selection – F# makes this easy:
|
1 2 3 4 |
let dividedBySelection=successfulQueries |> Array.partition (fun (x:SelectomeGene)-> x.SelectionSignature) let showsSelection= (float32 (fst dividedBySelection).Length)/ (float32 successfulQueries.Length) |
Step 4: Download Multiple Sequence Alignments – In order to decide how conserved a protein is relative to other proteins, we can download the multiple sequence alignment for each protein in this set and compare it to a particular protein of interest. In Selectome, each protein comes with a masked and unmasked alignment for both proteins and DNA. These objects are available from the SelectomeGene class and are lazily downloaded when requested from the Selectome server. These sequence alignment downloads are also cached for 30 days in a temporary directory to avoid waiting for downloads if you want to reload your script of interest. Once downloaded they are converted to full-fledged .NET BIO multiple sequence alignments, meaning one can do nearly anything with them. The example below gets the DNA alignment and the BLOSUM90 alignment score for the masked amino acid alignments.
|
1 2 3 4 |
let maskedAlignments = successfulQueries |> Array.map (fun x-> x.MaskedDNAAlignment) let alignmentScores= successfulQueries |> Array.map (fun x-> x.GetMaskedBlosum90AlignmentScore()) |
Step 5: Download the Gene Trees – The selectome gene defines a class,
SelectomeTree, that provides a great set of functions to query all the interesting metadata provided by selectome. These features are most usefully investigated by using the autocomplete functionality of your editor, but there is a lot of useful information! Some examples are also shown below.
|
1 2 3 4 5 6 7 8 |
let example=successfulQueries.[0] let tree=example.VertebrateTree let taxa=tree.TaxaPresent let selectedNode=tree.SelectedNodes.[0] selectedNode.BootStrap selectedNode.PValue selectedNode.Name ... |
Tree queries are also cached locally to avoid going back to the server in the event of repeated requests.
Step 6: Plot distribution of interest using the R data provider – You can call R plotting functions directly from F# using the R type provider. More information is available from that site, but the code snippet below is sufficient to produce a histogram of alignment scores, no need to dump to a flat file first!
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 |
#r @"C:\Users\Nigel\packages\RDotNet.FSharp.0.1.2.1\lib\net40\RDotNet.FSharp.dll" #r @"C:\Users\Nigel\packages\R.NET.1.5.5\lib\net40\RDotNet.dll" #r @"C:\Users\Nigel\packages\RProvider.1.0.1\lib\RProvider.dll" open RDotNet open RProvider open RProvider.``base`` open RProvider.graphics open RProvider.stats let args = new System.Collections.Generic.Dictionary<string,Object>() args.["x"] <- AlignmentScores args.["main"] <-"BLOSUM90 Alignment Scores" args.["col"] <-"blue" args.["breaks"]<-20 args.["xlab"]<-"BLOSUM90 Score" R.hist(args) |

Mono.Simd and the Mandlebrot Set.
C# and .NET are some of the fastest high level languages, but still cannot truly compete with C/C++ for low level speed, and C# code can be anywhere from 20%-300% slower. This is despite the fact that the C# compiler often gets as much information about a method as the C compiler. It has been suggested that SSE/SIMD instructions in C# could overcome some of these issues. To evaluate this, I took a famous computational task, re-coded it using C# SIMD instructions, and evaluated the performance by looking at the execution time and how the emitted assembly code compared to the optimal assembly code.
Background
In 2009, Miguel de Icaza demonstrated a framework that allows C# to use SSE2 intrinsic operations.This is now part of the Mono library and in theory, such methods can greatly save computational time (1.5X-16X), particularly for operations on byte arrays, a common task in bioinformatics.
Test Case
Computing the mandelbrot set is one of the tasks of the computer benchmarks game, which compares program speed in different languages. Currently, C# is listed as being slower than Java, though the programs in each language use different techniques and neither uses an optimal algorithm (see here for a better one). However, it makes a useful test case for raw computing speed. The challenge is to compute the picture shown below by recursively squaring a complex number and adding a constant to it.

The Algorithm
Complex numbers are a particularly challenging case because their multiplication is not a simple operation, but involves a rather annoyingly squaring of different terms and then adding/subtracting them.
These are not easy vector operations, and as will be shown later one result of this is that using SSE to speed up the values for one multiplication is useless (I tried, it was worse). So, the performance improvement comes from doing two elements of the loop at a time (this is how the current Java submission is faster than the C# one, though it does not use SIMD). The current C# version does the inner loop of the program without SIMD instructions as follows:
|
1 2 3 4 5 6 7 |
int i = 49; do { zi = zr * zi + zr * zi + ci; zr = tr - ti + cr; tr = zr * zr; ti = zi * zi; } while ((tr + ti <= 4.0) && (--i > 0)); |
This loop is iterating until convergence (<4) or until the max iterations (i<0) have expired. See the wikipedia page for a full explanation. Of interest here is that the “t” variables exist for pipelining purposes, so this is a reasonably optimized code snippet.
Trying to Implement the Optimal Solution Using Mono.Simd
Actually figuring out how to do the complex arthimetic with SSE2 is rather challenging. Fortunately Intel published a document giving the best solution as hand coded assembly, which involves using some special SSE3 instructions I was not aware of. Notably, the Intel optimized code is far faster than even their C code, but is only about 25% better than their optimized assembly code without SSE3.
The code below shows my attempt to implement the Intel approach in Mono. It should be pretty fast, however it may suffer from one fatal flaw. The comparison at the end to check for the final conditions currently requires unpacking both values in the Vector2D (when either Length.X or Length.Y have been < 4.0 at least once, then the loop stops). The comparison for both X and Y can be done in one operation in SIMD using the built in less than statement. However, I do not know how to turn that into a control loop function, as this requires a movmskps assembly function which mono does not seem to expose.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
int i = 49; int b = 0; do { Vector2d Zri=Zr*Zi; //calculate r*i for both Zi=Zri+Zri+Ci; //double that and add a constant Zr=Temp; //pre-calculated on previous loop var V0=Zr.InterleaveLow(Zi); //r0,i0 var V1=Zr.InterleaveHigh(Zi); //r1,i1 V0=V0*V0; //r0^2,i0^2 V1=V1*V1; var Length=V0.HorizontalAdd(V1); //(r0^2+i0^2),(r1^2+i1^2) Temp=V0.HorizontalSub(V1)+Cr; //(r0^2-i0^2),(r1^2-i1^2) //now to determine end condition, if (Length.X > 4.0) {b|=2; if (b==3) break;} if (Length.Y>;4.0) {b|=1; if (b==3) break;}} while (--i > 0); |
Results – It’s Faster on Ubuntu
Shown above are the times for three different algorithms. The first is the original best submission on the website. My SIMD submission is shown on the bottom. Because the SIMD version does two cycles within each inner loop, as the Java submission does, I tested the Java submission converted to C# as well. Compared to the current submission this version shows a 81% improvement, but clearly much of that is simply from doing 2 cycles in one loop. However, even once this change is made, the SIMD instructions still give a performance boost.
The Assembly Generated is Still Not Optimal
Although the assembly generated did include lots of SSE commands, in general inspecting the assembly I noticed several things.
-
Never unpack the double array values! My first attempt tried to do the SSE2 steps with those instructions, and then unpack the single values as needed. However, this failed prettty miserably, as accessing the double seemed to involve a lot of stack to register movement.
-
All the XMM registers are not used. The optimal version uses all of them, the C# version uses only 0-3, and soften moves things to and from the stack. Not using registers optimally seems to be a common problem though with C#.
Conclusions
The SIMD version was indeed a fair bit faster, which is nice! However, in this case it was not a game-changer. Most importantly though, it was really easy to use, and I think I might incorporate the byte array operations at some point in future code. This also gave me an appreciation for assembly, which in contrast to what I had heard is easy to read and seems easy to optimize. I just submitted the code to the shoot-out, assuming it works there it should be up soon, and I would love for someone to show me how to fix the end statement.
References
| 1. | ↑ | tr + ti <= 4.0) && (--i > 0 |
Accessing dbSNP with C# and the .NET Platform
NCBI Entrez can be accessed with many different platforms (python, R, etc.) , but I find .NET one of the best because the static typing makes it easy to infer what all the datafields mean, and navigate the data with much greater ease.
Documentation is sparse for this task, but here is how to access NCBI from the .NET platform. The example steps/program show how to query dbSNP for information about particular ids.
- Open visual studio, start a new project and add two Service References to the project: http://eutils.ncbi.nlm.nih.gov/soap/v2.0/eutils.wsdl and http://eutils.ncbi.nlm.nih.gov/soap/v2.0/efetch_snp.wsdl files. Note that the efetch depends on the database used, in this case “snp” other databases have different ones. You should now have two references:

More information is available here on setting up visual studio: http://www.ncbi.nlm.nih.gov/books/NBK55694/
2. Next up grab the data as shown in the example. Each SNP has A LOT more data, which can be inspected in the IDE.
1: static void Main(string[] args)
2: { 3: string dbSNPid="28357684";
4: efetch.eFetchRequest re = new efetch.eFetchRequest();
5: re.id = dbSNPid;6: var serv=new efetch.eUtilsServiceSoapClient();
7: var exchange = serv.run_eFetch(re); 8: var snpData = exchange.ExchangeSet.Rs[0];9: object[] dataToReport=new object[] {
10: snpData.Het.value,
11: snpData.hgvs.First(), 12: snpData.PrimarySequence.First().accession, 13: };14: Console.WriteLine( String.Join("\t",dataToReport.Select(x=>x.ToString()).ToArray()));
15: Console.ReadLine(); 16: }The following links contain other useful information for consuming the entrez webservice in C#.
Setting up visual studio: http://www.ncbi.nlm.nih.gov/books/NBK55694/
Using efetch: http://www.ncbi.nlm.nih.gov/books/NBK55694/#csharp_msvs.Call_HYPERLINK__chaptercha_8
Forming queries: http://www.biostars.org/p/3436/
More information: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/soap/v2.0/DOC/esoap_help.html
Also note that the first query takes much longer than subsequent ones, for reasons unknown to me at present.
![image_thumb[1] image_thumb[1]](https://i0.wp.com/evolvedmicrobe.com/blogs/wp-content/uploads/2013/09/image_thumb11.png?resize=244%2C237)

